In R, you can extract factor variables from a dataframe using various methods.
Let's create a sample data frame called mydata having 4 variables (ID, Gender, Region, Grade).
# Create a sample data frame
mydata <- data.frame(
ID = 1:5,
Gender = c("Male", "Female", "Male", "Male", "Female"),
Region = c("North", "South", "East", "West", "North"),
Grade = factor(c("A", "B", "A", "C", "B"))
)
You can use the str() function to see the structure or data type of a data frame. As shown in the result below, the variable "Grade" is a factor variable.
str(mydata)
'data.frame': 5 obs. of 4 variables: $ ID : int 1 2 3 4 5 $ Gender: chr "Male" "Female" "Male" "Male" ... $ Region: chr "North" "South" "East" "West" ... $ Grade : Factor w/ 3 levels "A","B","C": 1 2 1 3 2
How to Extract all Factor Variables in R
In the dataframe named "mydata", we know we have a factor variable "Grade". When we have multiple variables in a dataframe, we don't know the name of the factor variables in advance.
Base R
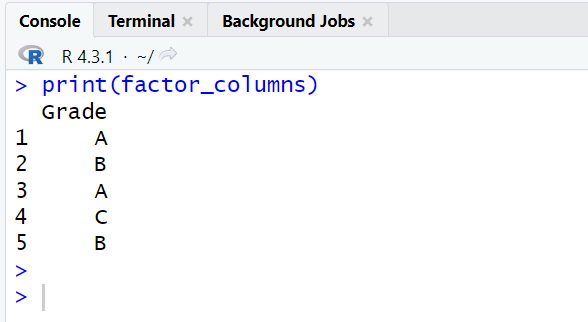
factor_columns <- mydata[sapply(mydata, is.factor)] print(factor_columns)
In base R, you can extract multiple factor columns (variables) using sapply function. The sapply function is a part of apply family of functions. They perform multiple iterations (loops) in R.
In dplyr package, the select_if function is used to select columns based on a condition. In this case, is.factor selects only the factor columns.
dplyr
library(dplyr) # Select factor columns using select_if() factor_columns <- mydata %>% select_if(is.factor) print(factor_columns)
Extract Factor Variables with more than 2 Unique Levels in R
Let's modify the "mydata" dataframe by adding one more factor variable for demonstration purpose.
mydata <- data.frame(
ID = 1:5,
Gender = c("Male", "Female", "Male", "Male", "Female"),
Region = factor(c("North", "South", "East", "West", "North")),
Grade = factor(c("A", "B", "A", "B", "B"))
)
Base R
In this code, we're using the sapply function to iterate through each column of the "mydata" data frame. For each column, we check if it's of factor data type (is.factor(col)) and if it has more than 2 unique levels (nlevels(col) > 2).
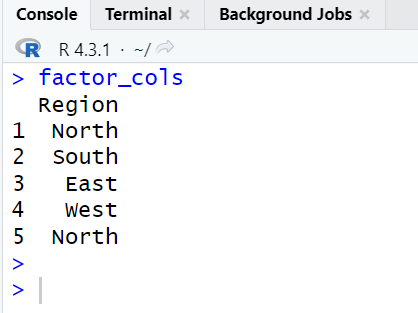
# Extract factor columns with more than 2 unique categories factor_cols0 <- sapply(mydata, function(col) is.factor(col) && nlevels(col) > 2) # Select columns based on the extracted factor column indicators factor_cols <- mydata[factor_cols0] print(factor_cols)
dplyr
In this code, we're using the dplyr package to work with data frames. The select_if function is used to select columns based on a condition. In this case, we're selecting columns that are of factor data type (is.factor(col)) and have more than 2 unique categories (nlevels(col) > 2).
library(dplyr) factor_cols <- mydata %>% select_if(function(col) is.factor(col) && nlevels(col) > 2) print(factor_cols)
Extracting Factor Variables with No Missing Values in R
Let's say you want to keep factor variables that have no missing values in R.
# Create a sample data frame
mydata <- data.frame(
name = c("Alice", "Bob", "Charlie", "Jon"),
city = c("Los Angeles", "New York", "Dallas", NA),
height = c(165.5, 180.0, 172.3, 181)
)
Base R
factor_cols <- sapply(mydata, is.factor) factor_no_missing <- colSums(is.na(mydata[factor_cols])) == 0 factor_no_missing_cols <- mydata[factor_cols] [factor_no_missing]
Let's see how the code works:
sapply(mydata, is.factor)checks whether each column in the dataframe is of type factor or not.colSums(is.na(mydata[factor_cols])) == 0checks if there are any missing values (NA) in the columns which are factors. It sums up the NAs in each factor column to find out non-missing values.mydata[factor_cols][factor_no_missing]selects only those columns that are factors and have no missing values based on the previous step.
dplyr
If you want to keep columns that have no missing values, you can use the select() function with where() in dplyr. select(where(is.factor)) selects only the factor columns. select(where(~ all(!is.na(.)))) selects columns where all values are not missing (NA).
library(dplyr) factor_no_missing_cols <- mydata %>% select(where(is.factor)) %>% select(where(~ all(!is.na(.))))


Share Share Tweet